Sequence
This Chapter describes window which is used for consecutive measurement of multiple samples. Open the window using the  icon or the Analysis - Sequence command from the
icon or the Analysis - Sequence command from the
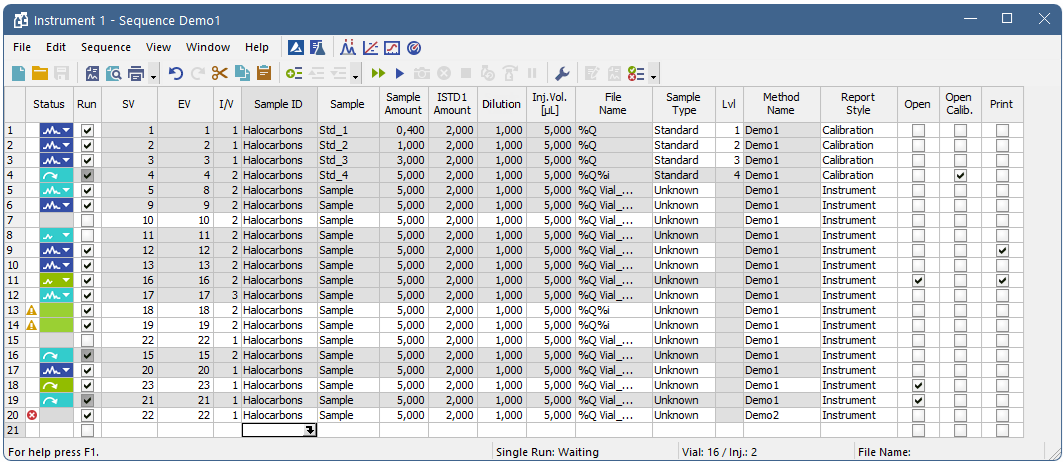
Sequence
The window includes a toolbar, sequence table and status bar.
Table description
Each line defines the settings for one or more injections. To process several vials from a single line enter a different value for the initial = start (SV) and last = end (EV) vial. The injection per vial I/V parameter indicates number of injections from one vial.
Status of the row in the sequence.
 Indicates a row with one or more errors, such row blocks the measurement of the sequence.
Indicates a row with one or more errors, such row blocks the measurement of the sequence.
 Indicates a row with one or more warnings, measurement will still be carried out.
Indicates a row with one or more warnings, measurement will still be carried out.
Point mouse cursor over the icon in the Status column to display a tooltip with detailed description about the injections - it displays how many were measured, skipped, aborted and how many are remaining on the specific row.
Note:
Analysis will be marked as finished after the analysis time elapsed. Note that the instrument method may be still running.
List of possible states of sequence rows
| Status | State | Description |
|---|---|---|

|
Ready | No analysis has been carried out from this row. Measurement will be carried out. |

|
Ready | No analysis has been carried out from this row. At least one vial has been skipped. Rest of the vials will be measured. |

|
Ready | Analysis has been stopped during acquisition. At least one chromatogram has been measured. Rest of the vials will be measured. |

|
Running | Analysis from this row is being carried out. |

|
Completed | All analyses on this row have been measured. |

|
Partially completed | Apart from the vial(s) where the Skip Vial command has been used, all analyses on this row have been measured. |

|
Partially completed | Command Skip Vial has been used on all vials. No more measurement will be carried out from this row. If icon changes color to green and back to original when (un)checking Run checkbox then there are vial(s) that still can be measured. |

|
Partially completed | Analysis has been stopped during acquisition. At least one chromatogram has been measured. Run checkbox is unckecked and measurement will not continue unless checking the Run checkbox. |

|
Disabled | Row that has been omitted from the sequence. Caused either by an error on the row or by unchecked Run checkbox. |
Note:
A text description will be printed in the printouts instead of colored tags in the Status column.
Already measured rows will be blocked against changes (dimmed) in all columns except for Run, Sample Type, Lvl, Report Style,
If there is a chromatogram resulting from a row, small arrow will appear in the icon - e.g.  (currently measured row with at least one already measured chromatogram). Left mouse click on the triangle will reveal options to open the chromatogram(s). See Fig "Open chromatograms in overlay".
(currently measured row with at least one already measured chromatogram). Left mouse click on the triangle will reveal options to open the chromatogram(s). See Fig "Open chromatograms in overlay".
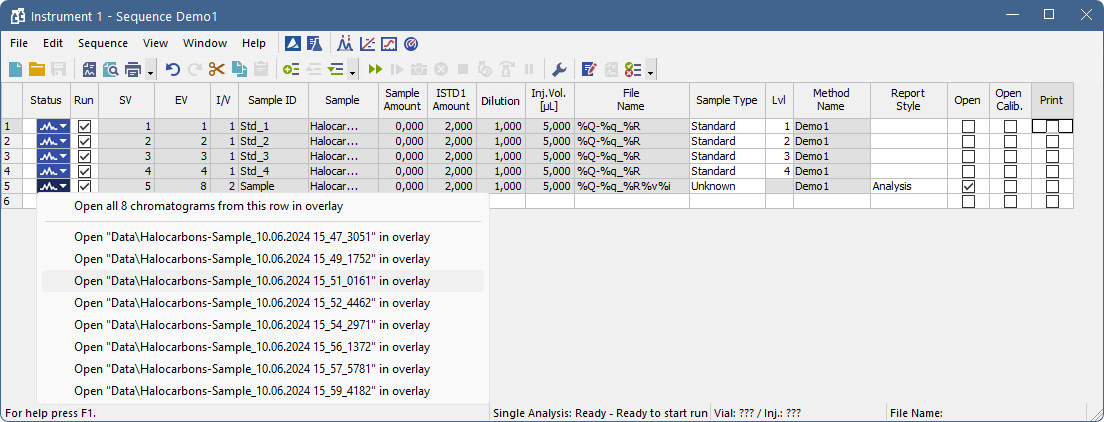
It is possible to open one specific chromatogram or open all chromatograms measured in this row in overlay.
Selected row(s) will return to their initial Status (all fields green) when the sequence is reset:
- By Run command, the
 icon or F5 or Ctrl + Q shortcut and clicking Reset button in the displayed dialog.
icon or F5 or Ctrl + Q shortcut and clicking Reset button in the displayed dialog. - Using the Edit - Reset Status command (Ctrl + E).
If the Sequence is reset, also its audit trail is reset. In GLP it is recommended to clone measured sequences using Save As option.
Selects whether the particular line in the Sequence Table will be measured. The column corresponds to the Run Lines field in Sequence Options dialog. Working with Run column is more suitable for discontinuous selection of lines or for exclusion of individual line from measurement.
Number of the first vial used in the processing of the given row.
Number of the last vial used in the processing of the given row. Its number must be the same or higher than the number entered in the SV field. All vials between SV and EV (these included) are measured.
If Sequence Options - Solve conflict of filename is set to Automatically (default value), then for more than one vial or injection, the File Name pattern is extended by %v and/or %i variables.
Note:
Some autosamplers may support alphanumerical or graphical setting of the SV or EV parameter. In such case, an arrow will appear in the right side of the given field, and clicking it will open particular Select Vial dialog.
Indicates how many injections will be processed from each vial based on this line in the Sequence Table.
If Sequence Options - Solve conflict of filename is set to Automatically (default value), then for more than one vial or injection, the File Name pattern is extended by %v and/or %i variables.
The sample identification field serves for information only. Maximum number of characters in this field is limited to 64. The parameter corresponds to the Sample ID field of the Single Analysis dialog. Sample ID is saved in the chromatogram and can be viewed or edited on Measurement Conditions - Instrument tab.
After clicking on the cell in this column the  icon will appear. Using it invokes the list where you can pick from the available variables for composing the name. Variables that can be used are described in the Chromatogram File Name section of the Single Analysis dialog
icon will appear. Using it invokes the list where you can pick from the available variables for composing the name. Variables that can be used are described in the Chromatogram File Name section of the Single Analysis dialog
The Sample description field serves for information only. Maximum number of characters in this field is limited to 64. The parameter corresponds to Sample field of the Single Analysis dialog. Sample is saved in the chromatogram and can be viewed or edited on Measurement Conditions - Instrument tab.
After clicking on the cell in this column the  icon will appear. Using it invokes the list where you can pick from the available variables for composing the name. Variables that can be used are described in the Chromatogram File Name section of the Single Analysis dialog.
icon will appear. Using it invokes the list where you can pick from the available variables for composing the name. Variables that can be used are described in the Chromatogram File Name section of the Single Analysis dialog.
Enable to add user comments. Text is editable directly in the cell or after clicking on dropdown arrow of the  icon in Comments dialog. Comments column is hidden and empty by default. Comments are saved in the chromatogram and can be viewed or edited on Measurement Conditions - Instrument tab.
icon in Comments dialog. Comments column is hidden and empty by default. Comments are saved in the chromatogram and can be viewed or edited on Measurement Conditions - Instrument tab.
The value is used in calibration calculations of individual components percentage in the ESTD and ISTD methods. In ISTD enter the sample amount without the addition. Enter values using the same units that were used in the calibration file (Units - Compound parameter in Calibration Options window). The default value is 0. The parameter corresponds to the Amount field of the Single Analysis dialog. Sample Amount is saved in the chromatogram and can be viewed or edited on Calculation - Common for all signals section of Results tab.
The amounts of the internal standards for the ISTD method. Other columns for entering amounts of ISTD2 - ISTD10 are hidden by default. This value has to be entered in the same units as those used in the calibration file (as defined in the Units - Compound field of the Calibration Options dialog). The default value is 0. The parameter corresponds to the ISTD1 Amount field of the Single Analysis dialog. ISTDs Amounts are saved in the chromatogram and can be viewed or edited on Calculation - Common for all signals section of Results tab.
This field holds the sample dilution factor - its value is used to multiply each calculated Amount in the Result Table. The default value is 1. The parameter corresponds to Dilution field in the Single Analysis dialog. Dilution is saved in the chromatogram and can be viewed or edited on Calculation - Common for all signals section of Results tab.
Defines the volume of injection. The response of all compounds in a calibration or when using calibrated calculations is modified by the ratio of this value and Default Injected Volume value from the calibration file. This parameter corresponds to the analogous field from Single Analysis dialog. The units are set in the Instrument Units dialog. Injection Volume is saved in the chromatogram and can be viewed or edited on Calculation - Common for all signals section of Results tab.
Caution:
When using a directly controlled autosampler this value will define the injection volume. The station then automatically verifies whether the entered volume corresponds to the current autosampler setting.
When the value set in this column is 0, no correction to the Default Injection Volume value will be made. However, when Inj.Vol. of 0 is entered into the sequence when using a directly controlled autosampler, the row of the sequence will be in error state.
AnalysisUserVar1 - AnalysisUserVar3
Sets numerical value of User Variables (up to 3 per row). User Variables can be used for User Column calculations. Columns AnalysisUserVar1 - AnalysisUserVar3 are hidden and set to value 0 by default. Name of the columns can be changed in Setup Columns... through Custom Name line. If the field is left empty, default name AnalysisUserVar1-AnalysisUserVar3 would remain filled in.
Caution:
Users who do no have the right to Edit Method in the User Accounts dialog cannot edit AnalysisUserVar1 - AnalysisUserVar3; fields will be grayed out.
Defines the chromatogram name pattern. The pattern may use a combination of fixed text and variables. The same variables may be used as in the Single Analysis dialog, plus %v and %i variables for numbering vials and injections when measuring multiple analysis from one line.
After clicking on the cell in this column the  icon will appear. Using it invokes the list where you can pick from the available variables for composing the file name. Variables are described in the Chromatogram File Name section of the Single Analysis dialog
icon will appear. Using it invokes the list where you can pick from the available variables for composing the file name. Variables are described in the Chromatogram File Name section of the Single Analysis dialog
It is possible to include also the name of a subfolder in the name to store chromatograms into subfolders of the Data Subdirectory. E.g. the MySubProject\MyFileName in the DEMO1 project will result in the MyFilename.prm stored in the C:\CLARITY\Datafiles\DEMO1\DATA\MySubproject folder. If the subfolder doesn't exist, it will be created when creating the chromatogram.
When the information on the sequence row are valid (the analysis from the row could proceed), the tooltip over the File Name cell will display the name(s) of the chromatogram file(s) (as it would look if the analysis started right away, with respect to time variables) and also the name of the calibration that will be created as a clone during the first analysis on that row (if calibration cloning is used and the row is the place where the cloning will occur).
Note:
If variable %n is being used, it will be independent from the same variable Single Analysis dialog.
Sets the type of the sample being measured. The possible options are:
Unknown - denotes an unknown sample.
Standard - specifies the calibration standard. Samples marked as Standard can have the calibration level filled in the Lvl column. Chromatograms marked as calibration standards in the time of creation will be stored in the calib subdirectory instead of the data subdirectory. If the Lvl is not filled, calibration (or recalibration) is not performed, chromatograms are only stored in the CALIB subdirectory.
Blank - in fact a calibration standard with no amount of added sample. The Blank is thus one point of a calibration curve (with the Amount of 0). Chromatograms marked as blanks in the time of creation will be stored in the calib subdirectory instead of the data subdirectory.
Bypass - allows for performing an analysis in the Active sequence with controlled autosampler without actually injecting a sample. This may be useful for system clean-up etc..
The row indicates an error and prevents running the sequence, if Standard (with filled Lvl) or Blank Sample Types are selected, and in the selected Method there is not defined the calibration file.
Caution:
In the case, the finishing of the analysis results in calibration (or recalibration) of calibration file, which is at the same time modified in the Calibration window, any unsaved changes are discarded, and the (re)calibration is performed.
Indicates the number of the calibration level that will be (re)calibrated. Cells in this columns are enabled only when Sample Type on the same row is set to Standard. If the Lvl is not filled, calibration (or recalibration) is not performed.
Indicates the name of the method governing all analyses set on a given line. If you point the mouse cursor to the cell, a tooltip will open, in which the name of the method including the entire path will be displayed.
Sets the report style to be used for printing. If printing is checked on the row, the report style has to be selected. After clicking on the cell in this column the  icon will appear. Using it opens the Select Report Style dialog. If you point the mouse cursor to the cell, a tooltip will open, in which the name of the report style including the entire path will be displayed.
icon will appear. Using it opens the Select Report Style dialog. If you point the mouse cursor to the cell, a tooltip will open, in which the name of the report style including the entire path will be displayed.
The commands Open, Open Calib., Print, Print to PDF, Export Data, Export AIA, Export TXT, Export EZChrom, Export Multidetector, Run Program, Program to Run, and Parameters generally correspond to analogous items in the Single Analysis - Post-run Options tab. All of these columns contain checkboxes, with the exception of the Program to Run column and the Parameters column.
Note:
When there is Print, Print to PDF or Export Data set in the Post-run Options, the Open chromatogram post-run option has to be selected too. Only chromatograms opened in the Chromatogram window can be reported. In sequence with either of the above functions set on a row without Open set as well, sequence row validation fails.
This checkbox allows user to set whether the particular chromatogram will be included in the SST calculations or not if opened in the chromatogram window.
This checkbox causes the created chromatogram to be opened with the latest stored version of the calibration file, as opposed to the linked calibration when not checked. The checkbox corresponds to the Open with stored calibration checkbox in the Calculation - Common for all signals section of Results tab.
This checkbox allows to close all chromatograms currently opened in Chromatogram window after the analysis is finished. If the checkbox Open is checked at the same time, the currently measured chromatogram will be opened in Chromatogram window. Other post-run operations always follow after Close all command.
This column serves for any information that belongs to the sequence only and is not transferred to the chromatograms.
Note:
To resize a column, point mouse cursor on the edge of the column to display  , left mouse click and drag to a desired width. All columns, apart from Status, can be resized.
, left mouse click and drag to a desired width. All columns, apart from Status, can be resized.
The status bar provides the following information:

State
If the sequence is running, the runtime of the actual sample is displayed. If the sequence is not running, status from the Instrument window in the form of <short status> - <long status> is displayed.
The vial number / injection number currently being measured.
The name of the chromatogram currently being measured.
Active/Pasive
Active/passive sequence flag.
Chromatogram name control flag showing the settings of the Solve conflict of filename section in the Sequence Options dialog